P04: Mapping the cis-regulatory landscape of the NKX2-1 locus and functional analyses of regulatory elements in thyroid development, disease, and evolution
Open postions: none
Principal investigator: Prof. Dr. Heiko Krude
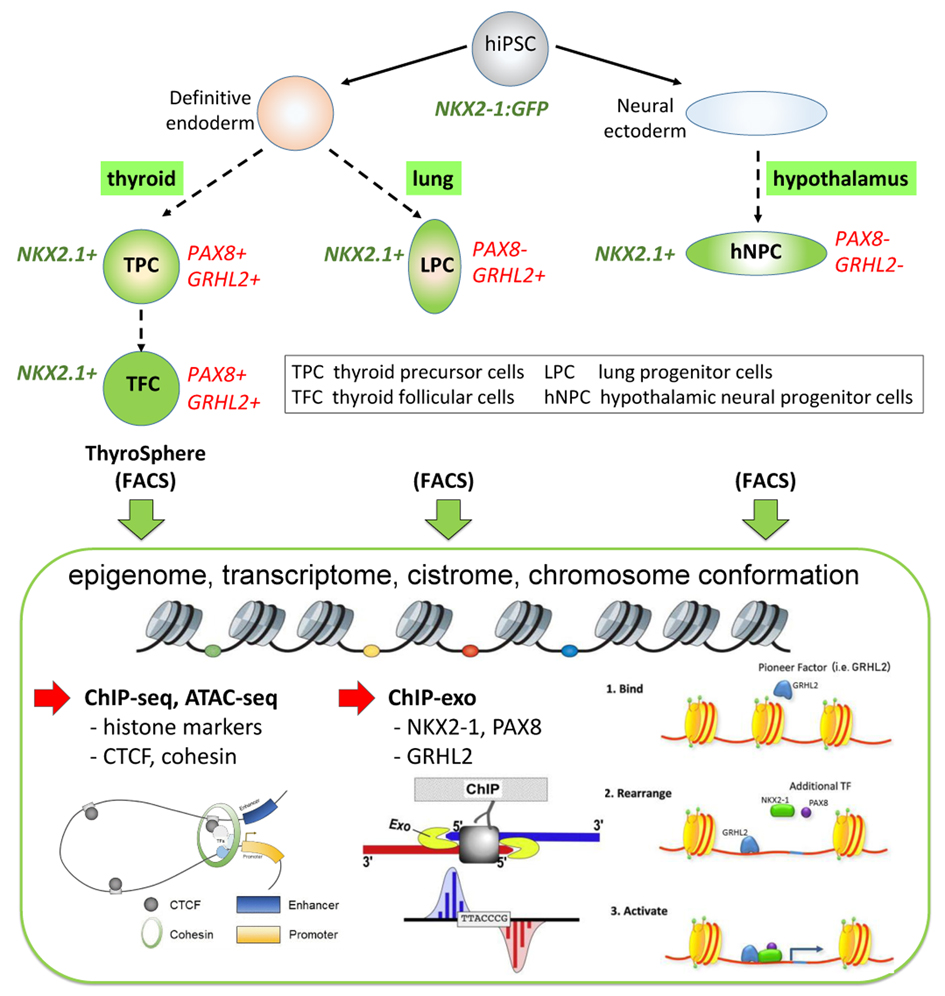 The molecular mechanisms causing congenital hypothyroidism due to thyroid dysgenesis (CHTD) remain largely unknown. Candidate gene approaches and whole exome analyses revealed disease-causing coding variants in less than 5% of all CHTD cases. We hypothesize that mutations or structural variants of non-coding regulatory elements are involved in the pathogenesis of CHTD. We will use whole genome sequencing to generate a novel genomic resource for a unique cohort of CHTD patients. These efforts will be accompanied by experimental studies with human induced pluripotent stem cells (hiPSC)-derived cell models aiming at a comprehensive thyroid-specific annotation of cis-regulatory elements (CREs) and functional genomics data for biologically relevant developmental stages. Although primary CHTD candidate genes, such as NKX2-1 and PAX8, are critical for the development of multiple tissues; the vast majority of CHTD cases present isolated thyroid phenotypes. To address emerging concepts of phenotypic modularity due to tissue-specific CRE mutations, we have devised a comparative experimental strategy comprising epigenomic and transcriptomic profiling of cell types representing the major NKX2-1-expressing cell lineages (thyroid, lung, and ventral forebrain) in the human embryo. In combination with in vivo functional genomic studies in zebrafish embryos, our experiments will provide a resource of developmentally relevant and tissue-specific genomic annotations to permit an accurate interpretation of genetic variation in the non-coding genome of CHTD patients. Our collaborators within the RU will be able to capitalize on our data to generate new hypotheses and models of TF-DNA interactions. Our data and analysis will shorten the diagnostic odyssey for CHTD patients and provide the broader scientific community with the data needed to further our understanding of the molecular mechanisms of thyroid diseases and the impact of epigenomic changes on cell fate decisions.
The molecular mechanisms causing congenital hypothyroidism due to thyroid dysgenesis (CHTD) remain largely unknown. Candidate gene approaches and whole exome analyses revealed disease-causing coding variants in less than 5% of all CHTD cases. We hypothesize that mutations or structural variants of non-coding regulatory elements are involved in the pathogenesis of CHTD. We will use whole genome sequencing to generate a novel genomic resource for a unique cohort of CHTD patients. These efforts will be accompanied by experimental studies with human induced pluripotent stem cells (hiPSC)-derived cell models aiming at a comprehensive thyroid-specific annotation of cis-regulatory elements (CREs) and functional genomics data for biologically relevant developmental stages. Although primary CHTD candidate genes, such as NKX2-1 and PAX8, are critical for the development of multiple tissues; the vast majority of CHTD cases present isolated thyroid phenotypes. To address emerging concepts of phenotypic modularity due to tissue-specific CRE mutations, we have devised a comparative experimental strategy comprising epigenomic and transcriptomic profiling of cell types representing the major NKX2-1-expressing cell lineages (thyroid, lung, and ventral forebrain) in the human embryo. In combination with in vivo functional genomic studies in zebrafish embryos, our experiments will provide a resource of developmentally relevant and tissue-specific genomic annotations to permit an accurate interpretation of genetic variation in the non-coding genome of CHTD patients. Our collaborators within the RU will be able to capitalize on our data to generate new hypotheses and models of TF-DNA interactions. Our data and analysis will shorten the diagnostic odyssey for CHTD patients and provide the broader scientific community with the data needed to further our understanding of the molecular mechanisms of thyroid diseases and the impact of epigenomic changes on cell fate decisions.